The R package netseer
netseer.Rmdnetseer

The goal of netseer is to predict the graph structure including new nodes and edges from a time series of graphs. The methodology is explained in the preprint (Kandanaarachchi et al. 2025). We will illustrate an example in this vignette.
Installation
You can install the development version of netseer from GitHub with:
# install.packages("devtools")
devtools::install_github("sevvandi/netseer")An example
This is a basic example which shows you how to predict a graph at the next time point. First let us generate some graphs.
library(netseer)
library(igraph)
#>
#> Attaching package: 'igraph'
#> The following objects are masked from 'package:stats':
#>
#> decompose, spectrum
#> The following object is masked from 'package:base':
#>
#> union
set.seed(2024)
edge_increase_val <- new_nodes_val <- del_edge_val <- 0.1
graphlist <- list()
graphlist[[1]] <- gr <- igraph::sample_pa(5, directed = FALSE)
for(i in 2:15){
gr <- generate_graph_exp(gr,
del_edge = del_edge_val,
new_nodes = new_nodes_val,
edge_increase = edge_increase_val )
graphlist[[i]] <- gr
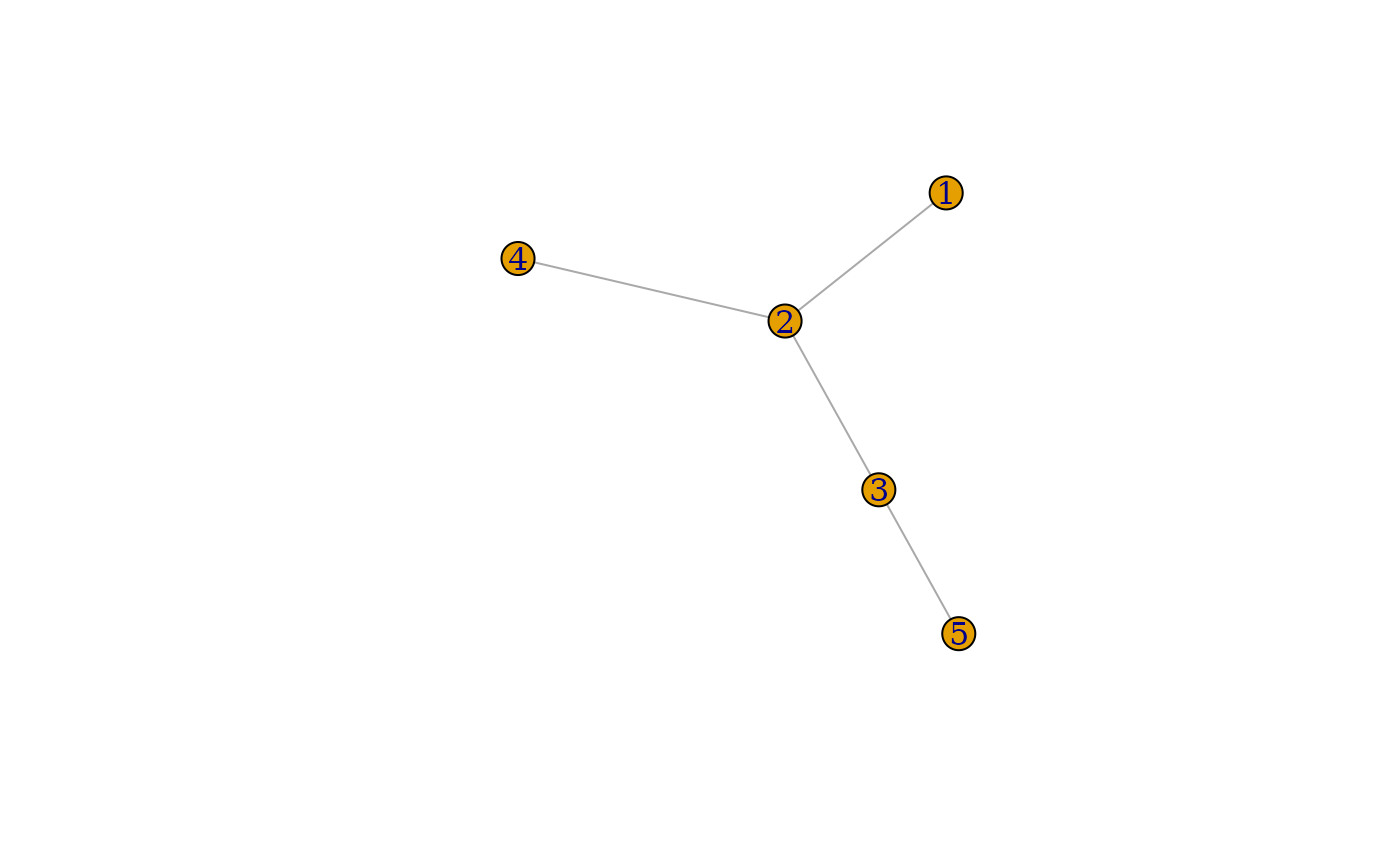
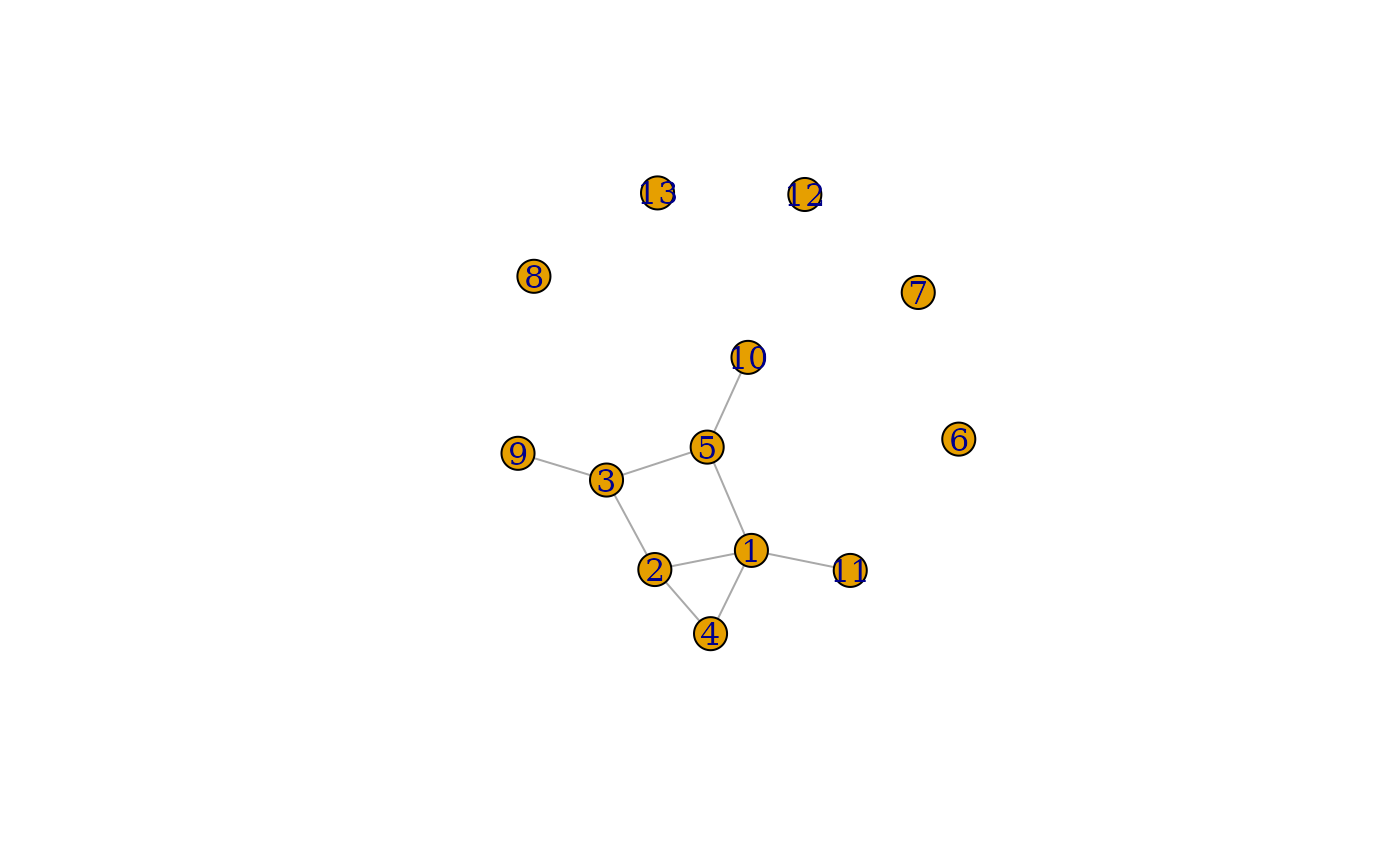
}The graphlist contains the list of graphs we generated. Each graph is an igraph object. Let’s plot a couple of them.
Predicting the next graph
Let’s predict the next graph. The argument specifies we want to predict the graph at the next time point.
grpred <- predict_graph(graphlist[1:15],h = 1)
#> Warning: 3 errors (1 unique) encountered for arima
#> [3] missing value where TRUE/FALSE needed
grpred
#> $graph_mean
#> IGRAPH d8e5b65 U--- 34 23 --
#> + edges from d8e5b65:
#> [1] 1-- 2 1-- 4 1-- 5 1-- 6 1--14 1--25 2-- 3 2-- 4 2--21 3-- 6
#> [11] 3-- 9 3--17 3--29 5--10 5--12 9--20 10--11 11--19 11--20 17--19
#> [21] 28--29 1-- 3 17--34
#>
#> $graph_lower
#> NULL
#>
#> $graph_upper
#> NULL
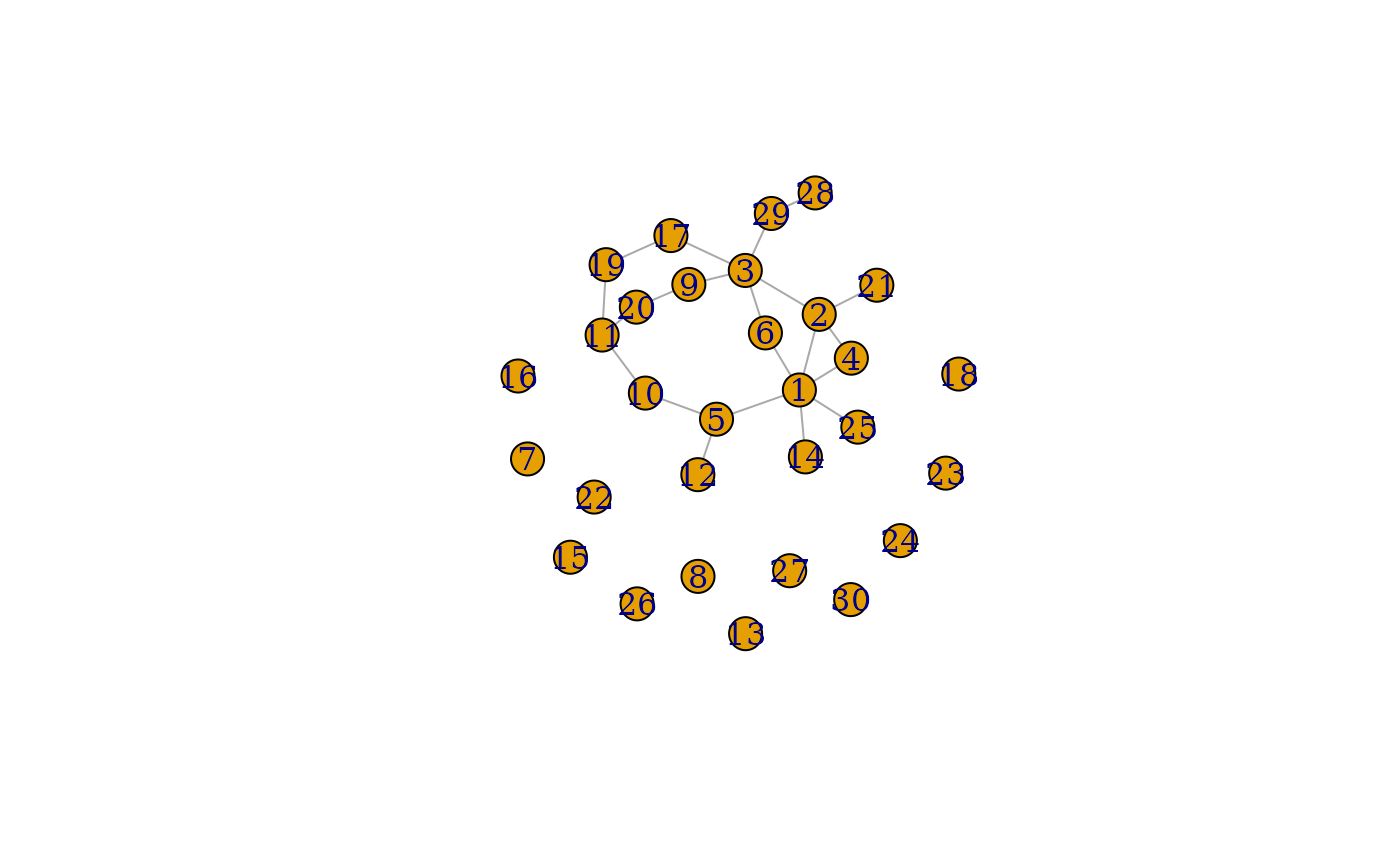
plot(grpred$graph_mean)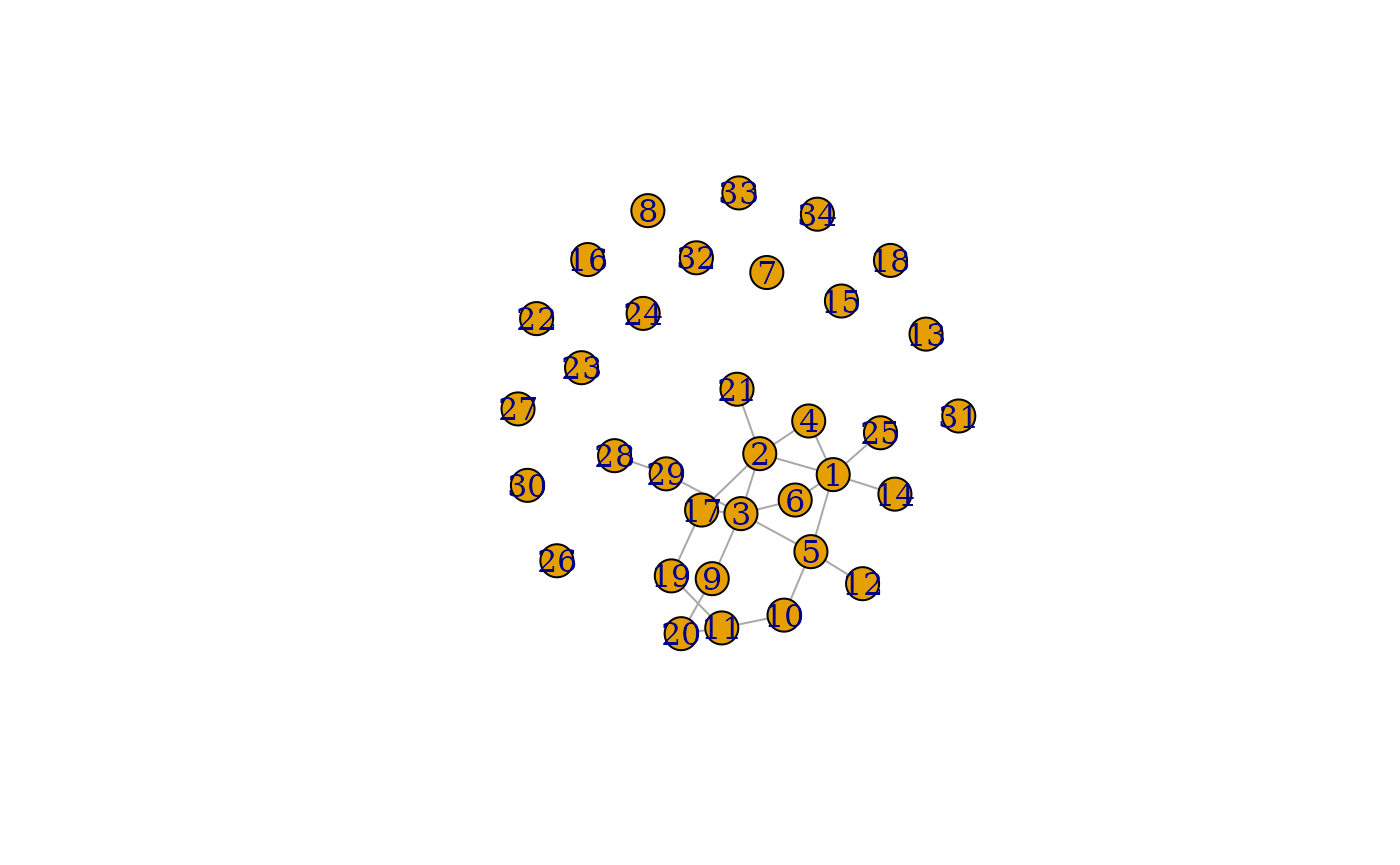
Predicting the graph at 2 time steps ahead
Now let us predict the graph at 2 time steps ahead with .
grpred2 <- predict_graph(graphlist[1:15], h = 2)
#> Warning: 3 errors (1 unique) encountered for arima
#> [3] missing value where TRUE/FALSE needed
grpred2
#> $graph_mean
#> IGRAPH 242163f U--- 37 26 --
#> + edges from 242163f:
#> [1] 1-- 2 1-- 4 1-- 5 1-- 6 1--14 1--25 2-- 3 2-- 4 2--21 3-- 6
#> [11] 3-- 9 3--17 3--29 5--10 5--12 9--20 10--11 11--19 11--20 17--19
#> [21] 28--29 1-- 3 3-- 4 2-- 5 4-- 5 17--37
#>
#> $graph_lower
#> NULL
#>
#> $graph_upper
#> NULL
plot(grpred2$graph_mean)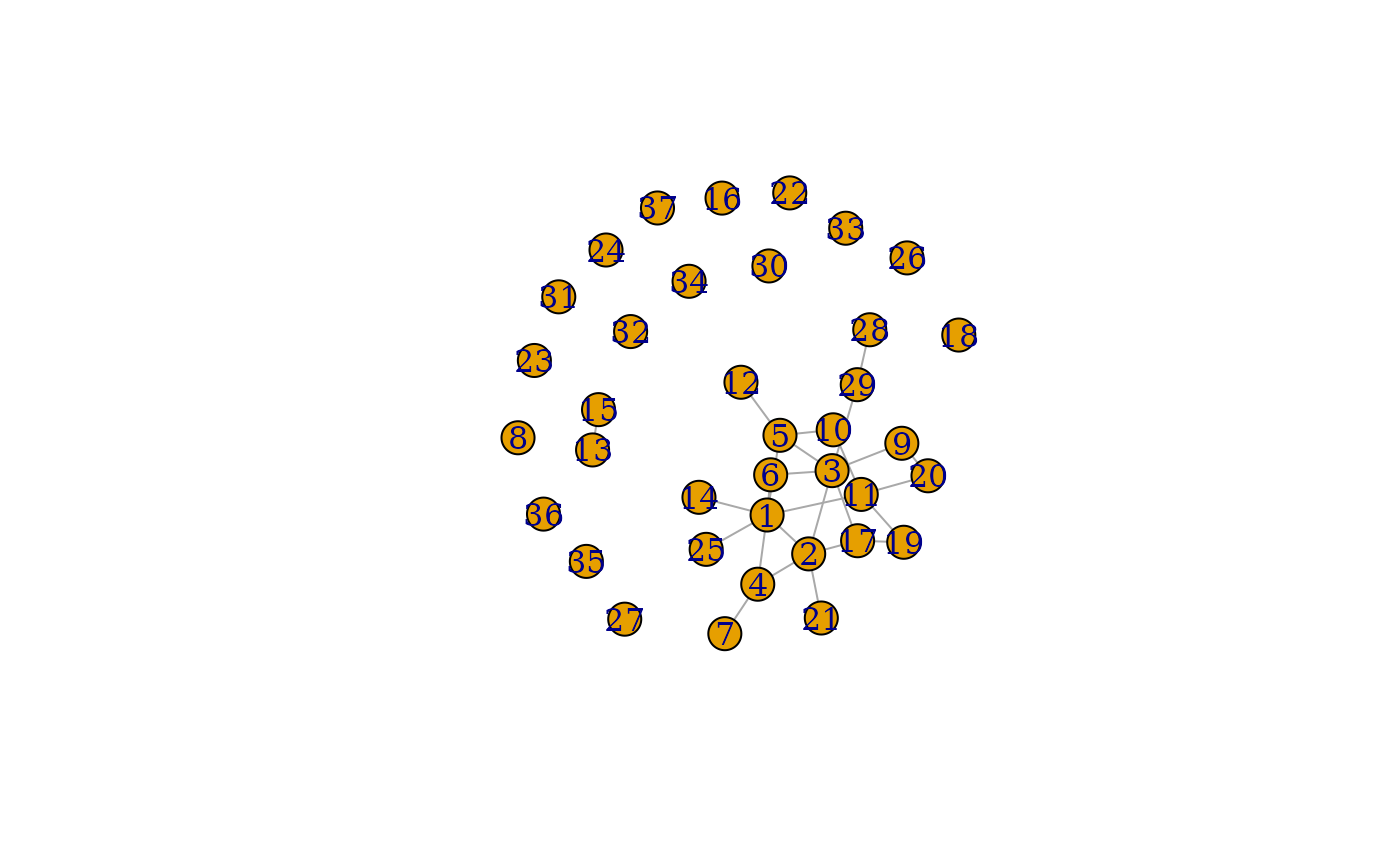
We see the predicted graph at has more vertices and edges than the graph at .
Predicting the graph at 3 time steps ahead
Similarly, we can predict the graph at 3 time steps ahead. We don’t have a limit on . But generally, as we get further into the future, the predictions are less accurate. This is with everything, not just graphs.
grpred3 <- predict_graph(graphlist[1:15], h = 3)
#> Warning: 3 errors (1 unique) encountered for arima
#> [3] missing value where TRUE/FALSE needed
grpred3
#> $graph_mean
#> IGRAPH 6d443be U--- 40 29 --
#> + edges from 6d443be:
#> [1] 1-- 2 1-- 4 1-- 5 1-- 6 1--14 1--25 2-- 3 2-- 4 2--21 3-- 6
#> [11] 3-- 9 3--17 3--29 5--10 5--12 9--20 10--11 11--19 11--20 17--19
#> [21] 28--29 1-- 3 3-- 4 2-- 5 4-- 6 5-- 6 2-- 9 1--10 17--40
#>
#> $graph_lower
#> NULL
#>
#> $graph_upper
#> NULL
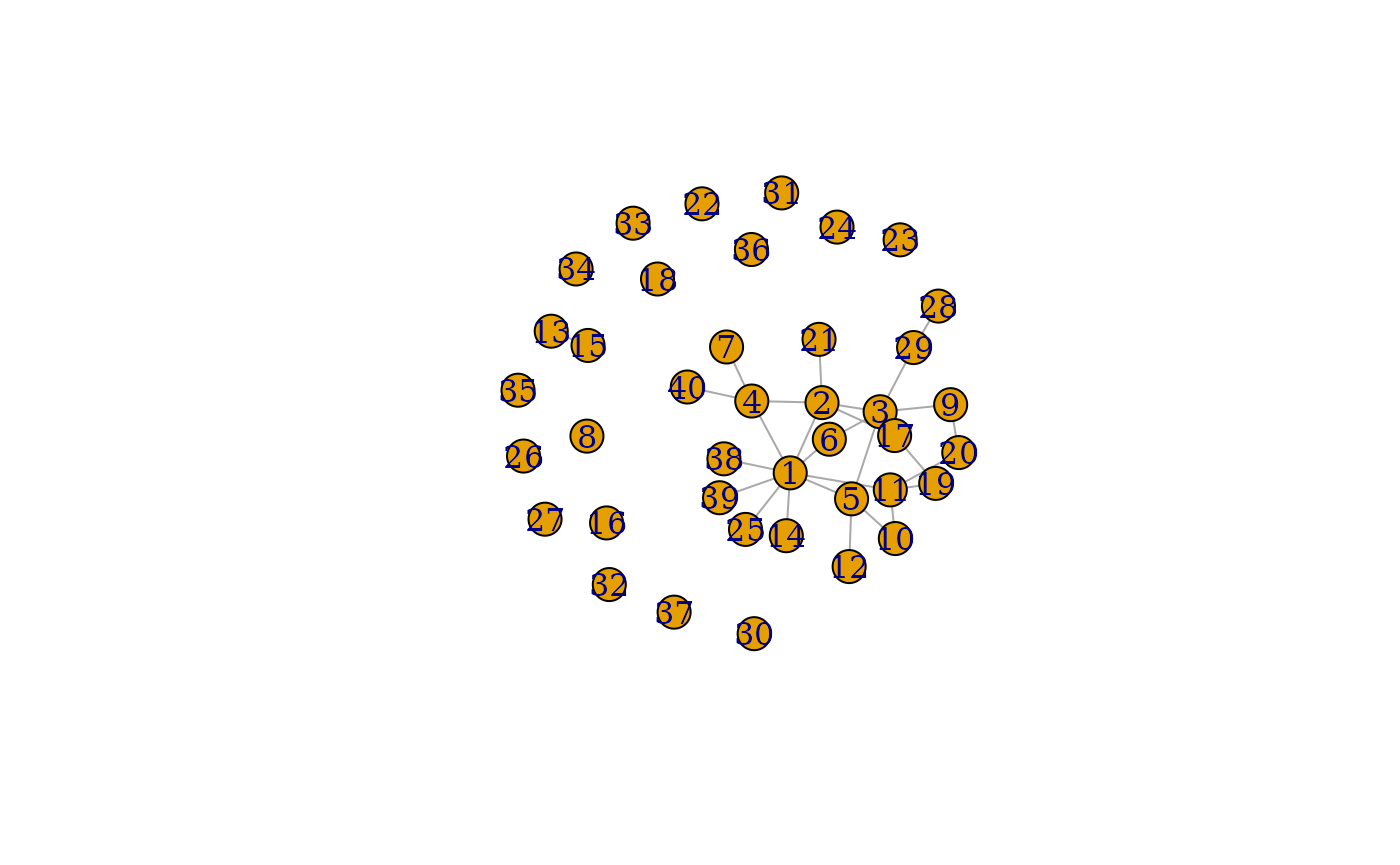
plot(grpred3$graph_mean)