Algorithm evaluation in other contexts
Revisiting pre-processing group fairness: A modular benchmarking framework
Authors: Brodie Oldfield, Ziqi Xu, Sevvandi KandanaarachchiVenue: ACM International Conference on Information and Knowledge Management, 2025
TLDR: FairPrep is a benchmarking framework that makes it easy to evaluate and compare pre‑processing fairness methods in machine learning using standardised, reproducible pipelines.
Revisiting pre-processing group fairness: A modular benchmarking framework
As machine learning systems become increasingly integrated into high-stakes decision-making processes, ensuring fairness in algorithmic outcomes has become a critical concern. Methods to mitigate bias typically fall into three categories: pre-processing, in-processing, and post-processing. While significant attention has been devoted to the latter two, pre-processing methods, which operate at the data level and offer advantages such as model-agnosticism and improved privacy compliance, have received comparatively less focus and lack standardised evaluation tools. In this work, we introduce FairPrep, an extensible and modular benchmarking framework designed to evaluate fairness-aware pre-processing techniques on tabular datasets. Built on the AIF360 platform, FairPrep allows seamless integration of datasets, fairness interventions, and predictive models. It features a batch-processing interface that enables efficient experimentation and automatic reporting of fairness and utility metrics. By offering standardised pipelines and supporting reproducible evaluations, FairPrep fills a critical gap in the fairness benchmarking landscape and provides a practical foundation for advancing data-level fairness research.
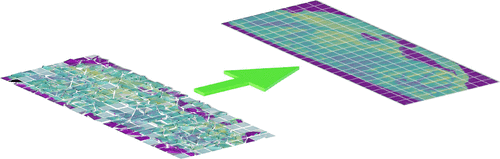
Comparison of tiling artifact removal methods in Secondary Ion Mass Spectrometry images
Authors: Sevvandi Kandanaarachchi, Wil Gardner, David L. J. Alexander, Benjamin W. Muir, Philippe A. Chouinard, Sheila G. Crewther, David J. Scurr, Mark Halliday, Paul J. PigramVenue: Analytical Chemistry, 2023
TLDR: There’s no one‑size‑fits‑all solution for removing tiling artifacts in large‑area ToF‑SIMS imaging—different correction methods work best for different datasets, so the choice of method should be data‑dependent.
Comparison of tiling artifact removal methods in Secondary Ion Mass Spectrometry images
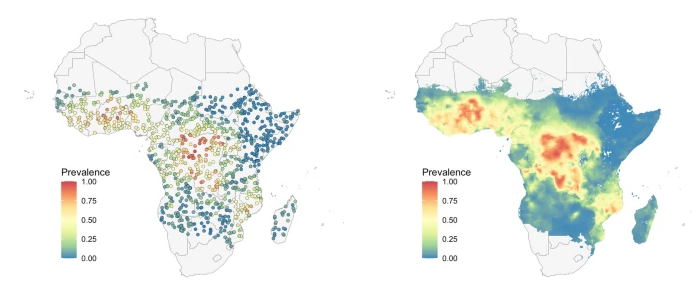
Comparison of new computational methods for spatial modelling of malaria
Authors: Spencer Wong, Jennifer A Flegg, Nick Golding, Sevvandi KandanaarachchiVenue: Malaria Journal, 2022
TLDR: An evaluation study of 4 geostatiscal methods for malaria modeling.